COVID1-9 News: A new study conducted by researchers from the Department of Immunobiology and the Department of Molecular Cellular and Developmental Biology at Yale School of Medicine-USA has found that the Omicron subvariants are evolving to further escape the human host immune system by...

www.thailandmedical.news
(fair use applies)
COVID-19 News: Yale Study Shows That Omicron Subvariants Are Evolving Further Through Mutations On ORF8 Proteins To Escape From MHC-I Recognition
Thailand Medical News
Jan 3 2023
COVID-19 News: Yale Study Shows That Omicron Subvariants Are Evolving Further Through Mutations On ORF8 Proteins To Escape From MHC-I Recognition
A new study conducted by researchers from the Department of Immunobiology and the Department of Molecular Cellular and Developmental Biology at Yale School of Medicine-USA has found that the Omicron subvariants are evolving to further escape the human host immune system by sprouting new mutations on the ORF8 proteins to help the virus escape from MHC-1 recognition.
Various past COVID-19 News coverages have shown that SARS-CoV-2 variants of concern (VOCs) possess mutations that confer resistance to neutralizing antibodies within the Spike protein and are associated with breakthrough infection and reinfection.
CD8+ T cell-mediated elimination of infected cells plays an important role in the antiviral adaptive immune response.
While it is known that numerous pathogenic viruses have developed strategies to evade host CD8+ T cell-mediated clearance not much is known about the escape from CD8+ T cell-mediated immunity by SARS-CoV-2 VOCs.
The study team demonstrated that all SARS-CoV-2 VOCs possess the ability to suppress MHC I expression.
The study team identified several viral genes that contribute to the suppression of MHC I expression. Notably, MHC-I upregulation was strongly inhibited after SARS-CoV-2 infection in vivo.
Although earlier VOCs possess similar capacity as the ancestral strain to suppress MHC I, Omicron subvariants exhibit a greater ability to suppress surface MHC-I expressions.
The study findings suggest that, in addition to escape from neutralizing antibodies, the success of Omicron subvariants to cause breakthrough infection and reinfection may in part be due to its optimized evasion from CD 8 T cell recognition.
The study findings were published on a preprint server and are currently being peer reviewed.
SARS-CoV-2 variants of concern (VOCs) possess mutations that confer resistance to neutralizing antibodies within the Spike protein and are associated with breakthrough infection and reinfection. By contrast, less is known about the escape from CD8+ T cell-mediated immunity by VOC. Here, we...

www.biorxiv.org
Increasing breakthrough infection and reinfection events are associated with the emergence of various new Omicron variants and sub-lineages. Breakthrough infections and reinfections are likely driven by significant increases in transmissibility, evasion from innate immunity, and escape from neutralization by vaccine/infection-induced antibodies.
It has been noted that the outstanding features of the Omicron variant are the considerably enhanced escape from the antibody neutralization and increased infectivity than the earlier VOCs, due to its heavily mutated Spike protein.
Though the Omicron variant and its subvariants harbor a far greater number of mutations in its genome compared to those in previous VOCs, T cell epitopes remain generally intact.
The severe acute respiratory distress syndrome coronavirus 2 (SARS-CoV-2) Omicron variant is spreading rapidly, even in vaccinated individuals, raising concerns about immune escape. Here, we studied neutralizing antibodies and T cell responses targeting SARS-CoV-2 D614G [wild type (WT)] and the...

pubmed.ncbi.nlm.nih.gov
Typically, the CD8+ cytotoxic T lymphocyte (CTL) recognizes and kills infected cells and eliminates the source of replicating viruses. Antigen presentation by major histocompatibility complex class I (MHC-I) is a critical step for the activation of antigen-specific CD8+ T c ells and the subsequent killing of infected cells. Viral peptides processed by the cellular proteasome complex are loaded on MHC-I molecule in the endoplasmic reticulum and translocate to the cell surface to be recognized by antigen-specific CD8+ T cells. To successfully establish infection and replicate in the host, many viruses have acquired the ability to inhibit MHC-I processing and presentation of viral antigens.
Similarly, the SARS-CoV-2 virus utilizes its viral proteins to interfere with the MHC-I pathway.
SARS-CoV-2 ORF8 protein induces autophagic degradation of MHC-I and confers resistance to CTL surveillance.
COVID-19, caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), has become a global pandemic and has claimed over 2 million lives worldwide. Although the genetic sequences of SARS-CoV and SARS-CoV-2 have high homology, the clinical and pathological characteristics of COVID-19...

pubmed.ncbi.nlm.nih.gov
Research from the first 3 months of the pandemic showed a rapid evolution of the SARS-CoV-2 ORF8 gene including isolates with 382nt deletion spanning ORF7b-ORF8 gene region, which is associated with robust T cell response and milder clinical outcome.
In this study, we analyzed full-length SARS-CoV-2 genomes from multiple countries to determine early trends in the evolutionary dynamics of the novel COVID-19 pandemic. Results indicated SARS-CoV-2 evolved early into at least three phylogenetic groups, characterized by positive selection at...

pubmed.ncbi.nlm.nih.gov
To date, limited genetic changes in the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) genome have been described. Here, we report a 382-nucleotide (nt) deletion in SARS-CoV-2 that truncates open reading frame 7b (ORF7b) and ORF8, removing the ORF8 transcription regulatory sequence...

pubmed.ncbi.nlm.nih.gov
National Medical Research Council Singapore.

pubmed.ncbi.nlm.nih.gov
The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) variants of concern (VOCs) that have become dominant as the pandemic progresses bear the ORF8 mutation together with multiple spike mutations. A 382-nucleotide deletion (Δ382) in the ORF7b and ORF8 regions has been associated with...

pubmed.ncbi.nlm.nih.gov
All these previous findings collectively raised a question of whether Omicron and its ORF8 protein have evolved to further enhance the ability to shut down MHC-I, thereby evading from antigen-specific memory CD8+ T cells established by previous infection or vaccination.
The study team performed a systematic analysis of the capacity of SARS-CoV-2variants to downregulate MHC-I presentation. Our data demonstrated vigorous suppression of MHC-I surface expression by the ancestral SARS-CoV-2 and minimal evolution in modulating MHC-I pathway by earlier VOCs.
The findings remarkably showed that the latest Omicron subvariants have acquired an enhanced ability in modulating MHC-I pathway.
The study uncovered the intrinsically potent ability of SARS-CoV-2 to shut down the host MHC-I system by using live, authentic SARS-CoV-2 variants as well as the functional analysis of variant-specific mutations in ORF8 gene, a key viral protein for both MHC-I evasion and adaptation to the host.
The study further identified multiple other viral genes that confer redundant function in MHC I suppression.
The study team showed that respiratory epithelial cells infected in vivo with SARS-CoV-2 failed to upregulate MHC-I, whereas those infected with influenza virus robustly elevated MHC-I expression.
Importantly the study findings revealed that the most recent Omicron subvariants have superior capacity to suppress MHC-I compared to the earlier isolates.
The study findings demonstrated that most variants of concern/interest possess unique mutations within ORF8 gene. However, none of these ORF8 mutations led to further reduction of MHC-I in cells expressing these molecules. Notably, the Omicron variant and its descendants lacked in non-synonymous mutations in their ORF8 gene, indicating that the mechanisms for superior MHC-I suppression by the Omicron sub-lineages must be located outside the ORF8 protein.
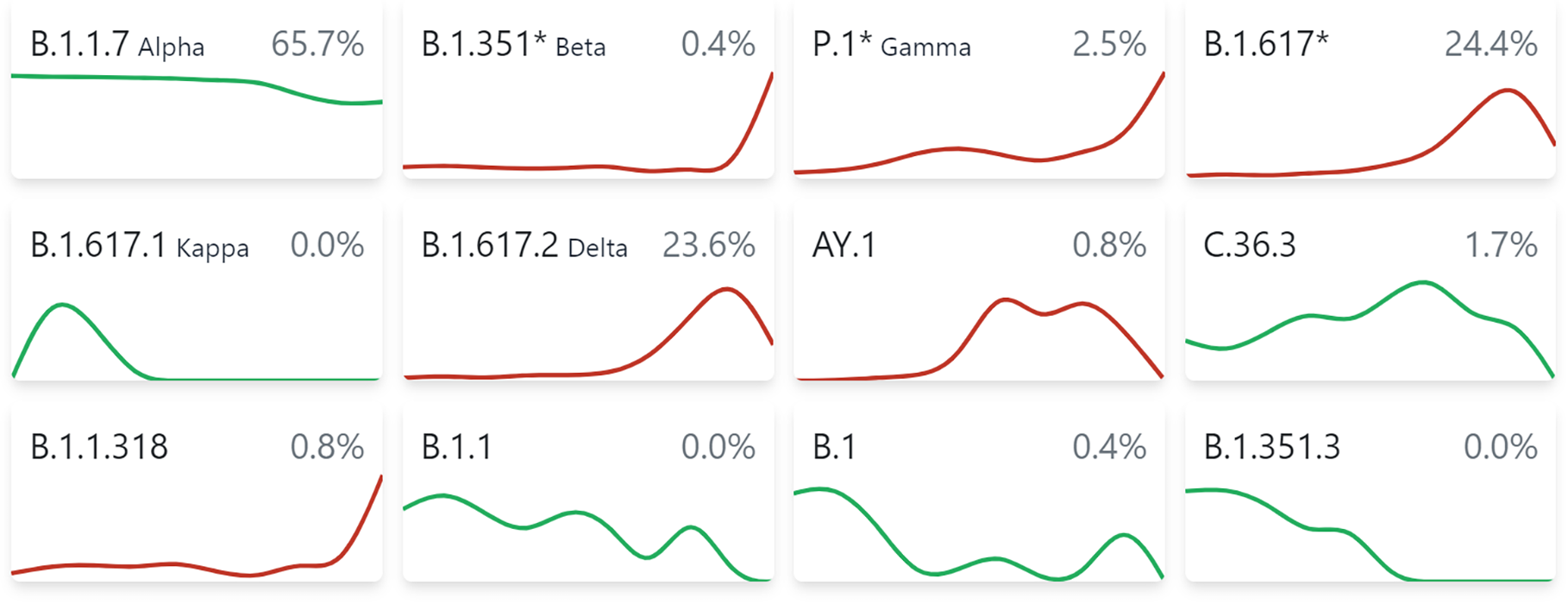
The study data showed that the ORF8 mutation in the B.1.1.7/Alpha lineage abrogated its function in MHC I modulation. Given the truncation mutation likely rendering ORF8 in the Alpha variant non-functional, this raises a question as to how such mutations might be tolerated.
Multiple functions beyond MHC-I downregulation are documented for SARS-CoV-2 ORF8, which include inhibition of type I IFN, ISGs, or NF-kB signaling, epigenetic modulation through histone mimicry and induction of proinflammatory cytokines from macrophages and monocytes via IL-17RA.
Numerous studies interestingly showed that SARS-CoV-2 ORF8 is actively secreted into the cell culture media in a signal peptide-dependent manner when it is overexpressed in vitro.
There is a rising global concern for the recently emerged novel coronavirus (2019-nCoV). Full genomic sequences have been released by the worldwide scientific community in the last few weeks to understand the evolutionary origin and molecular characteristics of this virus. Taking advantage of...

pubmed.ncbi.nlm.nih.gov
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is still adapting to its new human host. Attention has focussed on the viral spike protein, but substantial variation has been seen in the ORF8 gene. Here, we show that SARS-CoV-2 ORF8 protein undergoes signal peptide-mediated...

www.biorxiv.org
An accurate diagnostic test for early severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection is the key weapon to control the coronavirus disease 2019 (COVID-19) pandemic. We previously reported that the SARS-CoV-2 genome contains a unique orf8 accessory gene absent from other...

pubmed.ncbi.nlm.nih.gov
Also, ORF8 peptides and anti-ORF8 antibodies can be detected abundantly in serum of patients, suggesting the relevance of the active secretion of ORF8 to actual infection in humans.
An accurate diagnostic test for early severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection is the key weapon to control the coronavirus disease 2019 (COVID-19) pandemic. We previously reported that the SARS-CoV-2 genome contains a unique orf8 accessory gene absent from other...

pubmed.ncbi.nlm.nih.gov
The Alpha variant likely acquired compensatory mechanisms that enabled its successful transmission until the next variant came along. ORF8 is implicated in adaptation to the human host during the SARS-CoV outbreak and it is known that ORF8 is the hypervariable genomic region among the SARS-CoV and bat SARS related coronaviruses.
Studies from early in the COVID-19 pandemic observed the variability and rapid evolution of SARS-CoV-2 ORF8 gene. Notably, SARS-CoV-2 isolates with 382nt deletion spanning ORF7b-ORF8 gene region were observed in Singapore which correlated with robust T cell response and mild clinical outcome.
Hence, mutations in ORF8 gene may play a key role in modulating viral pathogenesis and adaptation to the host by regulating MHC-I levels and ISGs.
The enhanced immune evasion by VOCs has been well documented for escape from neutralizing antibodies and from innate immune responses.
The study findings demonstrated that the ability to reduce MHC-I expression remained unchanged throughout the pre-Omicron VOC evolution.
These study findings suggested three important perspectives on the MHC-I evasion strategy of SARSCoV-2:
1) SARS-CoV-2 utilizes multiple redundant strategies to suppress MHC-I expression. For example, considering B.1.1.7 retained an intact ability to shut down MHC-I, the impaired MHC-I evasion by B.1.1.7 ORF8 is likely compensated by the redundant and/or compensatory functions of other viral proteins including E, M, and ORF7a. In addition, B.1.1.7 lineage has been shown to express an increased sub-genomic RNA and protein abundance of ORF6, which suppresses MHC-I at the transcriptional level by interfering with STAT1- IRF1-NLRC5 axis. The multi-tiered MHC-I evasion mechanisms thus work redundantly to ensure escape from CTL killing.
2) MHC-I downregulation may not only impair CTL recognition of infected cells for killing but may also impair priming of CD8 T cells. Indeed, the frequency of circulating SARS-CoV-2 specific memory CD8+ T cells in SARS-CoV-2 infected individuals are ~10 fold lower than for influenza or Epstein-Barr virus-specific T cell populations, which indicates the suboptimal induction of memory CD8+ T cells following SARS-CoV-2 infection in human.
3) Given that the variants of concern had not further evolved to downregulate MHC-I more strongly than the original strain except for the Omicron subvariants, SARS-CoV-2 ancestral virus was already fully equipped to escape from CD8+ T cell-mediated immunity with respect to downregulation of MHC-I expression and is under less evolutionary pressure to further optimize the evasion strategy than those from type I IFNs or antibodies.
However, mutations and evasion from particular HLA-restricted CTL epitopes have been observed in circulating SARS-CoV-2 and VOCs.
Genome-wide screening of epitopes suggested the CD8+ T and CD4+ T cell epitopes are broadly distributed throughout SARS-CoV-2 genome, and the estimated numbers of epitopes per individual are at least 17 for CD8+ T and 19 for CD4+ T cells, respectively, and thus functional T cell evasion by VOCs is very limited.
This implies that MHC-I downregulation may be a more efficient way for viruses to avoid CTL surveillance than introducing mutations in epitopes.
The importance of MHC-I evasion by SARS-CoV-2 is also highlighted by the fact that no genetic mutations or variations in the MHC-I pathway has thus far been identified as a risk factor for severe COVID, unlike innate immune pathways involving TLRs and type I IFNs.
SARS-CoV-2 infection in both human and preclinical models have shown to induce antigen-specific CD8+ T cell responses and the early CTL response correlated with a milder disease outcome in human.
The magnitude of SARS-CoV-2-specific T cell responses correlates inversely with human disease severity, suggesting T cell involvement in primary control. Whereas many COVID-19 vaccines focus on establishing humoral immunity to viral spike protein, vaccine-elicited T cell immunity may bolster...

pubmed.ncbi.nlm.nih.gov
Over the past year, numerous studies in the peer reviewed and preprint literature have reported on the virological, epidemiological and clinical characteristics of the coronavirus, SARS-CoV-2. To date, 25 studies have investigated and identified SARS-CoV-2-derived T cell epitopes in humans...

pubmed.ncbi.nlm.nih.gov
Virus-specific humoral and cellular immunity act synergistically to protect the host from viral infection. We interrogate the dynamic changes of virological and immunological parameters in 12 patients with symptomatic acute SARS-CoV-2 infection from disease onset to convalescence or death. We...

pubmed.ncbi.nlm.nih.gov
Adoptive transfer of serum or IgG from convalescent animals alone, however, is enough to reduce viral load in recipients after SARS-CoV-2 challenge in mice and nonhuman primates and neutralizing antibody is shown to be a strong correlate of protection.
The protective roles of CD8+ T cell-mediated immunity appear to be more important in the absence of the optimal humoral responses/neutralizing antibody.
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has caused more than 160 million infections and more than 3 million deaths worldwide. Although effective vaccines are currently being deployed, the adaptive immune determinants that promote viral clearance and confer protection remain...

pubmed.ncbi.nlm.nih.gov
Patients with cancer have high mortality from coronavirus disease 2019 (COVID-19), and the immune parameters that dictate clinical outcomes remain unknown. In a cohort of 100 patients with cancer who were hospitalized for COVID-19, patients with hematologic cancer had higher mortality relative...

pubmed.ncbi.nlm.nih.gov
Blood anti-ORF8 antibodies can be used as the highly sensitive clinical marker for SARS-CoV-2 infection early (~14 days) after symptom onset, which suggests the role of ORF8 in the very early stage of the disease.
An accurate diagnostic test for early severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection is the key weapon to control the coronavirus disease 2019 (COVID-19) pandemic. We previously reported that the SARS-CoV-2 genome contains a unique orf8 accessory gene absent from other...

pubmed.ncbi.nlm.nih.gov
The SARS-CoV-2 virus emerged in December 2019 and has caused a worldwide pandemic due to the lack of any pre-existing immunity. Accurate serology testing is urgently needed to help diagnose infection, determine past exposure of populations and assess the response to a future vaccine. The...

pubmed.ncbi.nlm.nih.gov
Hence, ORF8-mediated MHC-I downregulation can therefore precede antigen presentation and hinder priming of viral antigen-specific CD8+ T cell immune responses.
Robust MHC-I shutdown by SARS-CoV-2 may explain in part the less effective protection by CD8+ T cells and the less impact of CD8+ T cell absence compared with humoral immunity.
The study findings shed light on the intrinsically potent ability of SARSCoV-2 to avoid the MHC-I mediated antigen presentation to CD8+ T cells. Importantly, the study findings showed a complete inhibition of MHC-I upregulation in lung epithelial cells infected with SARS-CoV-2 at the early stage of infection in a mouse model. Since the ability of ORF8 to downregulate MHC-I is a newly acquired feature in SARS-CoV-2 ORF8 and is absent in SARS-CoV ORF8, it is possible that ORF8 played a role in the efficient replication and transmission of SARS-CoV-2 in human and contributed to its pandemic potential.
The study findings provide insights into SARS-CoV-2 pathogenesis and evolution and predicts difficulty for CD8 T cell-based therapeutic approaches to COVID-19.
 m.timesofindia.com
m.timesofindia.com